Pipeline
Pipelines define the processing of data within PDAL. They describe how point cloud data are read, processed and written. PDAL internally constructs a pipeline to perform data translation operations using translate, for example. While specific applications are useful in many contexts, a pipeline provides useful advantages for many workflows:
You have a record of the operation(s) applied to the data
You can construct a skeleton of an operation and substitute specific options (filenames, for example)
You can construct complex operations using the JSON manipulation facilities of whatever language you want.
Note
pipeline is used to invoke pipeline operations via the command line.
Introduction
A PDAL processing pipeline is represented in JSON. The structure may either:
a JSON object, with a key called
pipelinewhose value is an array of inferred or explicit PDAL Stage Objects representations.a JSON array, being the array described above without being encapsulated by a JSON object.
Simple Example
A simple PDAL pipeline, inferring the appropriate drivers for the reader and writer from filenames, and able to be specified as a set of sequential steps:
[
"input.las",
{
"type":"filters.crop",
"bounds":"([0,100],[0,100])"
},
"output.bpf"
]

A simple pipeline to convert LAS to BPF while only keeping points inside the box \([0 \leq x \leq 100, 0 \leq y \leq 100]\).
Reprojection Example
A more complex PDAL pipeline reprojects the stage tagged A1, merges
the result with B, and writes the merged output to a GeoTIFF file
with the writers.gdal writer:
[
{
"filename":"A.las",
"spatialreference":"EPSG:26916"
},
{
"type":"filters.reprojection",
"in_srs":"EPSG:26916",
"out_srs":"EPSG:4326",
"tag":"A2"
},
{
"filename":"B.las",
"tag":"B"
},
{
"type":"filters.merge",
"tag":"merged",
"inputs":[
"A2",
"B"
]
},
{
"type":"writers.gdal",
"filename":"output.tif"
}
]
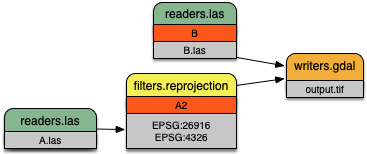
A more complex pipeline that merges two inputs together but uses
filters.reprojection to transform the coordinate system of
file B.las from UTM to Geographic.
Point Views and Multiple Outputs
Some filters produce sets of points as output. filters.splitter, for example, creates a point set for each tile (rectangular area) in which input points exist. Each of these output sets is called a point view. Point views are carried through a PDAL pipeline individually. Some writers can produce separate output for each input point view. These writers use a placeholder character (#) in the output filename which is replaced by an incrementing integer for each input point view.
The following pipeline provides an example of writing multiple output files from a single pipeline. The crop filter creates two output point views (one for each specified geometry) and the writer creates output files ‘output1.las’ and ‘output2.las’ containing the two sets of points:
[
"input.las",
{
"type" : "filters.crop",
"bounds" : [ "([0, 75], [0, 75])", "([50, 125], [50, 125])" ]
},
"output#.las"
]
Processing Modes
PDAL process data in one of two ways: standard mode or stream mode. With standard mode, all input is read into memory before it is processed. Many algorithms require standard mode processing because they need access to all points. Operations that do sorting or require neighbors of points, for example, require access to all points.
For operations that don’t require access to all points, PDAL provides stream mode. Stream mode processes points through a pipeline in chunks, which reduces memory requirements.
When using pdal translate or
pdal pipeline
PDAL uses stream mode if possible. If stream mode can’t be used
the applications fall back to standard mode processing. Streamable stages are
tagged in the stage documentation with a blue bar. Users can explicitly
choose to use standard mode by using the --nostream option. Users of the PDAL API can explicitly control the selection of the PDAL
processing mode.
Pipelines
Pipeline Array
PDAL JSON pipelines are an array of stages.
Note
In versions of PDAL prior to 1.9, the array of stages needed to be the value of a key named “pipeline” which was encapsulated in an object. The earlier format is still accepted for backward compatibility.
Old format:
{
"pipeline" :
[
"inputfile",
"outputfile"
]
}
Equivalent new format:
[
"inputfile",
"outputfile"
]
The pipeline array may have any number of string or Stage Objects elements.
String elements shall be interpreted as filenames. PDAL will attempt to infer the proper driver from the file extension and position in the array. A writer stage will only be created if the string is the final element in the array.
Stage Objects
For more on PDAL stages and their options, check the PDAL documentation on Readers, Writers, and Filters.
A stage object may have a member with the name
tagwhose value is a string. The purpose of the tag is to cross-reference this stage within other stages. Eachtagmust be unique.A stage object may have a member with the name
inputswhose value is an array of strings. Each element in the array is the tag of another stage to be set as input to the current stage.Stages are processed sequentially in the order listed. An empty default input list is created when interpretation of the pipeline begins.
Reader stages will disregard the
inputsmember. When the current stage is a reader it is added to the default input list.If
inputsis specified for a writer or filter, those inputs are used for the current stage. The default input list is replaced with the current stage.If
inputsis not specified for a writer or filter, the default input list is used for the current stage. The default input list is replaced with the current stage.A
tagmentioned in another stage’sinputsmust have been previously defined in thepipelinearray.A reader or writer stage object may have a member with the name
typewhose value is a string. Thetypemust specify a valid PDAL reader or writer name.A filter stage object must have a member with the name
typewhose value is a string. Thetypemust specify a valid PDAL filter name.A stage object may have additional members with names corresponding to stage-specific option names and their respective values. Values provided as JSON objects or arrays will be stringified and parsed within the stage. Some options allow multiple inputs. In those cases, provide the option values as a JSON array.
A
user_dataoption can be added to any stage object and it will be carried through to any serialized pipeline output.All stages support the
option_fileoption that allows options to be places in a separate file. See Option Files for details.
Filename Globbing
A filename may contain the wildcard character
*to match any string of characters. This can be useful if working with multiple input files in a directory (e.g., merging all files).Filename globbing ONLY works in pipeline file specifications. It doesn’t work when a filename is provided as an option through a command-line application like
pdal pipelineorpdal translate.
Option Files
All stages accept the option file option that allows extra options for a
stage to be placed in a separate file. The value of the option is the filename
in which the additional options are located.
Option files can be written using either JSON syntax or command line syntax. When using the JSON syntax, the format is a block of options just as if the options were placed in a pipeline:
{
"minor_version": 4,
"out_srs": "EPSG_4326"
}
When using the command line syntax, the options are specified as they would be on the command line without the need to qualify the option names with the stage name:
--minor_version=4 --out_srs="EPSG_4326"
Extended Examples
BPF to LAS
The following pipeline converts the input file from BPF to LAS, inferring both the reader and writer type, and setting a number of options on the writer stage.
[
"utm15.bpf",
{
"filename":"out2.las",
"scale_x":0.01,
"offset_x":311898.23,
"scale_y":0.01,
"offset_y":4703909.84,
"scale_z":0.01,
"offset_z":7.385474
}
]
Python HAG
In our next example, the reader and writer types are once again inferred. After reading the input file, the ferry filter is used to copy the Z dimension into a new height above ground (HAG) dimension. Next, the filters.python is used with a Python script to compute height above ground values by comparing the Z values to a surface model. These height above ground values are then written back into the Z dimension for further analysis. See the Python code at hag.py.
See also
filters.hag_nn describes using a specific filter to do this job in more detail.
[
"autzen.las",
{
"type":"filters.ferry",
"dimensions":"Z=>HAG"
},
{
"type":"filters.python",
"script":"hag.py",
"function":"filter",
"module":"anything"
},
"autzen-hag.las"
]
DTM
A common task is to create a digital terrain model (DTM) from the input point cloud. This pipeline infers the reader type, applies an approximate ground segmentation filter using filters.smrf, filters out all points but the ground returns (classification value of 2) using the filters.range, and then creates the DTM using the writers.gdal.
[
"autzen-full.las",
{
"type":"filters.smrf",
"window":33,
"slope":1.0,
"threshold":0.15,
"cell":1.0
},
{
"type":"filters.range",
"limits":"Classification[2:2]"
},
{
"type":"writers.gdal",
"filename":"autzen-surface.tif",
"output_type":"min",
"gdaldriver":"GTiff",
"window_size":3,
"resolution":1.0
}
]
Decimate & Colorize
This example still infers the reader and writer types while applying options on both. The pipeline decimates the input LAS file by keeping every other point, and then colorizes the points using the provided raster image. The output is written as ASCII text.
[
{
"filename":"1.2-with-color.las",
"spatialreference":"EPSG:2993"
},
{
"type":"filters.decimation",
"step":2,
"offset":1
},
{
"type":"filters.colorization",
"raster":"autzen.tif",
"dimensions": ["Red:1:1", "Green:2:1", "Blue:3:1" ]
},
{
"filename":"junk.txt",
"delimiter":",",
"write_header":false
}
]
Reproject
Our first example with multiple readers, this pipeline infers the reader types, and assigns spatial reference information to each. filters.reprojection filter reprojects data to the specified output spatial reference system.
[
{
"filename":"1.2-with-color.las",
"spatialreference":"EPSG:2027"
},
{
"filename":"1.2-with-color.las",
"spatialreference":"EPSG:2027"
},
{
"type":"filters.reprojection",
"out_srs":"EPSG:2028"
}
]
Globbed Inputs
Finally, we capture another merge pipeline demonstrating the ability to glob multiple input LAS files from a given directory.
[
"/path/to/data/\*.las",
"output.las"
]
See also
The PDAL source tree contains a number of example pipelines that are used for testing. You might find these inspiring. Go to https://github.com/PDAL/PDAL/tree/master/test/data/pipeline to find more.
Note
Issuing the command pdal info --options will list all available
stages and their options. See info for more.